Zalosnik et al. Sci Rep. 2021
CIQUIBIC-CONICET-UNC
Contact : +54 351 5353855

Zalosnik et al. Sci Rep. 2021

Nigra et al. Oxid Med Cell Longev. 2021

Alvarez Bolaño et al. Colloids SurfB 2021
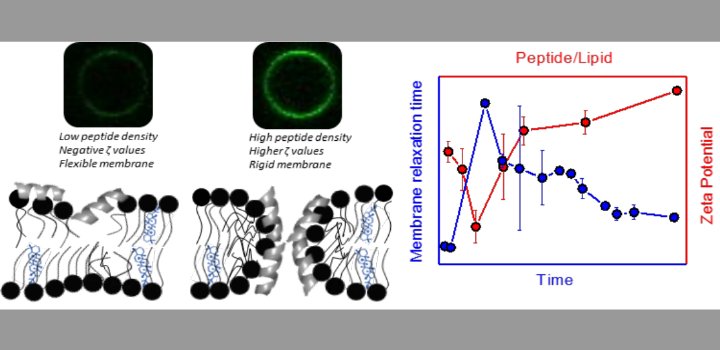
Alvares et al. BBA Advances 2021

Nocelli NE et al. helion. 2021

Lardone and at. JBC 2021

Wagner et al. FASEB J. 2021
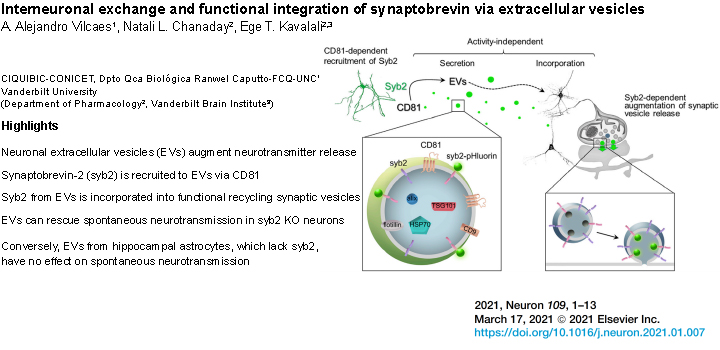
Vilcaes et al. J. Neuron. 2021

Cecchini NM et al. Plant J. 2020

Guido ME et al. 2020. Cell Mol Neurobiol

Prucca CG et al. Biochem J. 2020

Andrea G Albarracín Orio et al. Commun Biol. 2020

Zulueta Díaz AND BBA-Biomem. 2020
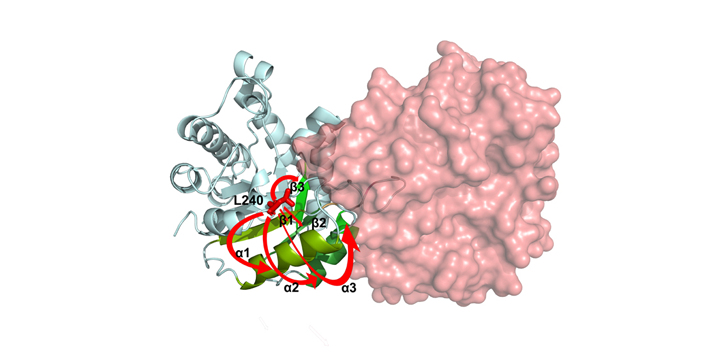
Romero J. Arch Biochem Biophys. 2020

Smith et al. Plant Physiol Biochem. 2020

Rodríguez-Berdini et al. J Biol Chem. 2020

Colque CA et al. Antimicrob Agents Chemother. 2020

Vilcaes et al. Int J Mol Sci. 2020

Cardozo Gizzi et al. Nat Protoc. 2020
Simultaneous observation of 3D chromatin organization and transcription at the single-cell level and with high spatial resolution may hold the key to unveiling the mechanisms regulating embryonic development, cell differentiation and even disease. We recently developed Hi-M, a technology that enables the sequential labeling, 3D imaging and localization of multiple genomic DNA loci, together with RNA expression, in single cells within whole, intact Drosophila embryos. Importantly, Hi-M enables simultaneous detection of RNA expression and chromosome organization without requiring sample unmounting and primary probe rehybridization. Here, we provide a step-by-step protocol describing the design of probes, the preparation of samples, the stable immobilization of embryos in microfluidic chambers, and the complete procedure for image acquisition. The combined RNA/DNA fluorescence in situ hybridization procedure takes 4–5 d, including embryo collection. In addition, we describe image analysis software to segment nuclei, detect genomic spots, correct for drift and produce Hi-M matrices. A typical Hi-M experiment takes 1–2 d to complete all rounds of labeling and imaging and 4 additional days for image analysis. This technology can be easily expanded to investigate cell differentiation in cultured cells or organization of chromatin within complex tissues.
Authors: Cardozo Gizzi AM, Espinola SM, Gurgo J, Houbron C, JB plug, DI Cats, Nollmann M.

Ruggiero et al. Immunol Cell Biol. 2020

Cardozo Gizzi A. et al. J Mol Biol. 2020

Chumpen S. et al. Biosci Rep. 2020
Malcolm M. et al. Front Cell Neurosci. 2019
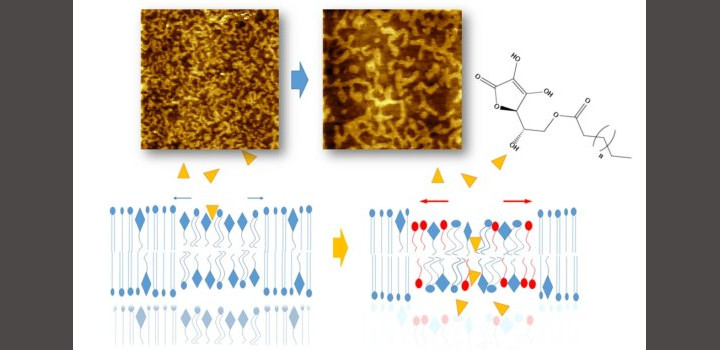
Zulueta Y. et al. Colloids Surf B Biointerfaces. 2020

Curtins JA et al. Biochem J. 2019

Lic. Antonella Colque will make a stay in Denmark
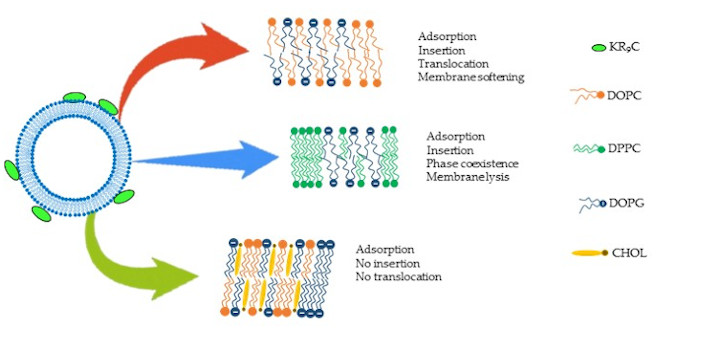
Crossio M. et al. Biomolecules. 2019
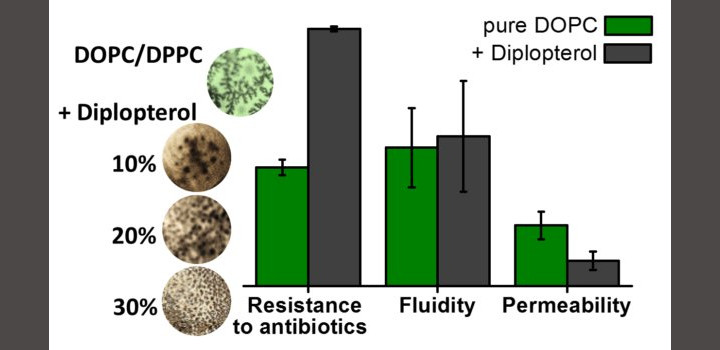
A Mangiarotti. et al. BBA-Biomembr. 2019

Monday 16December 2019 to12:30hs– Integrative Building Auditorium. FCQ.

Fellow of CIQUIBIC honored for her work presented at SAIB 2019
Rios M. et al. Front Cell Neurosci. 2019

By Dr. Horacio de la Iglesia (University of Washington)

Thursday 19of Septemberfrom 2019to11:00hs– Integrative Building Auditorium. FCQ.

Bertoldi ML et al. Front Cell Neurosci. 2019

Monday 9 of September from 2019 to 12:00hs– Integrative Building Auditorium. Fcq.

Quiroga & Valdez Meth. Mol. Biol. 2019